How will FAIR principles change literature review?
Making research more FAIR to increase transparency, improve quality, and rebuild public trust in science

Integration of FAIR - findable, accessible, interoperable and reproducible - principles into literature review comes at a critical time. Academic publication is expanding at an unprecedented pace and traditional research methods are badly in need of an upgrade to keep up. Public skepticism of previously trustworthy outlets (media, government, etc.) makes the transparency enabled by FAIR critical for building trust in academic review.
It is no longer possible for researchers to read all of the publications in their field. Even relatively specialized fields like "medical device stent" generate thousands of publications per year. Unfortunately, this means that most scientists are only exposed to a small fraction of the available research, and usually in a very biased way. When researchers use search engines to find relevant data they may look at only the top few results until they find something fitting for their own research. Systematic reviews, which we cover in another post, are a way to combat this problem, but another problem remains.
When any kind of review is done, the process behind that review remains inscrutable. Which articles did the researcher consider? Why did they discard some articles and keep others? Online review applications make high resolution review referencing possible and FAIR data practices add transparency and accountability.
Finding data representing a literature review is difficult . You can find publications discussing review results, but typically can't access the actual review data and project information. Existing review platforms aren't built around data sharing. Sysrev is the first platform to offer open access data to public projects. This makes it possible to find reviews of important topics. For example, you can now google "gene hunter sysrev" and find a project reviewing genes in thousands of abstracts.
When programmers try to solve problems they (like everybody else) usually rely on google first. Because of the large number of open source repositories on github, code can be quickly found to solve an enormous number of problems. Findable, open source code is critical in the development of many programming projects.
With thousands of literature reviews on the same topic (just search prospero for reviews on 'lead'), similar tools are in great need for the world of systematic review.
Accessible data will reduce redundancy in systematic review
Of course, it is not enough to find relevant existing research, we must be able to leverage that research as well. Actually, this is an even more critical problem in systematic review.
A publication is one of the primary results of most research. In the past decade journals have become more serious about sharing data related to research. Tools like figshare, NCBI's gene expression omnibus, and many more provide hosting services for different kinds of publication data.
In a similar manner, when reviews are published they could contain direct links to detailed information on the entire review process. This would enable others to use the generated review data to fact check reviews, extend reviews, answer new questions, or create new reviews.
On sysrev.com, public projects make all of the data in a systematic review open to the public. Simply visit the articles page, filter the data as desired, and click the export button. Programming packages are also in development to allow computational access (github.com/sysrev) have also been developed to allow computational access.
For instance, you could export all the data from the 40 person, 57000 article, massive NIEHS Hallmark and Key Characteristic Mapping Project at sysrev.com/p/3588/articles. Or download all the gene annotations from 10,000 sentences curated in the gene hunter review at sysrev.com/p/3144/articles.
Accessible systematic review data will reduce redundancy in the systematic review space. Searching for lead on prospero (a registry for systematic reviews) yields 4185 registered reviews. Certainly many of these reviews involve similar, sometimes identical data extraction steps.
If you're wondering why all of this is so important then you probably aren't a systematic reviewer. These reviews can take years and have serious ramifications for processes like creation of public policy and defining best practices for medical treatments. The next time you see a statistic about drug efficacy consider whether it came from a review of many related clinical trials or a single clinical trial.
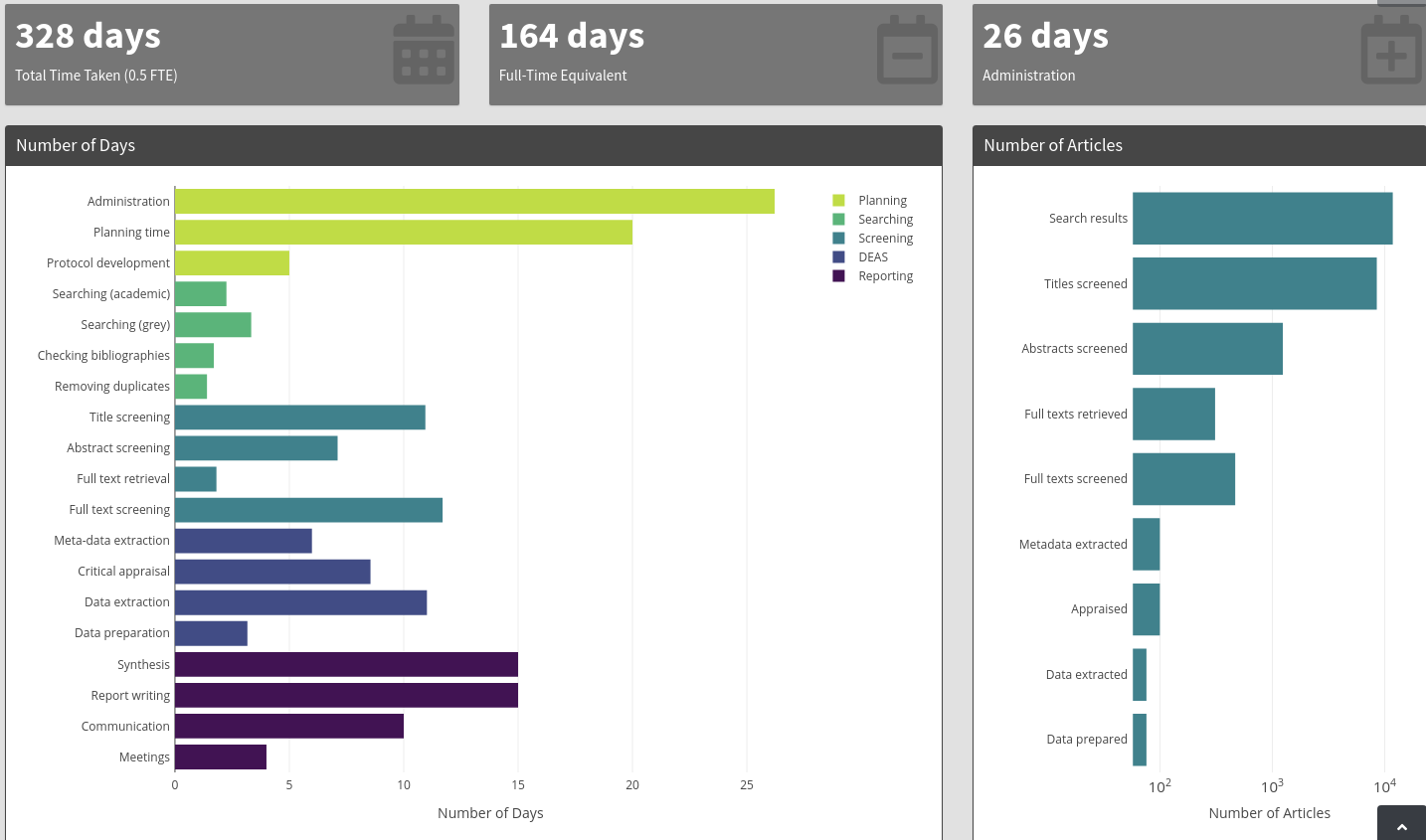
According to Neal Haddaway and Martin Westgate the average systematic review between 2012 and 2017 took 164 days of full time work. Their web application predictor.org breaks down the time for each step in the review process. High quality reviews take an astounding amount of time.
Interoperability and Reproducibility are focuses of sysrev.com development.
Sysrev offers Python/R (github.com/sysrev) packages for accessing and manipulating data at public reviews. Users can integrate public review data into their own projects or create new reviews with these packages. Integrations with new data sources and existing literature tools are also coming.
Sysrev also offers project cloning. Currently this feature can only be used by those who are familiar with programming, but soon sysrev will have a clone button on every systematic review.
curl -X POST \
'https://sysrev.com/web-api/clone-project?api-token=<token>&
project-id=3144&
new-project-name=cloneReview&
articles=true&
labels=true&
members=true&
answers=true' \Project cloning is a real super power for systematic reviews. Once cloning is fully available it will enable reviewers to more easily validate an existing review, or create a periodical reviews, or do things we haven't even considered. These principles will make systematic review much more transparent and accountable.
Without FAIR principles it is possible for special interests to capture the review process. Cochrane reviews are created by thousands of volunteers and trusted by the global community. However, in September 2018, Nature reported that "Governing board of the evidence-based medicine group may now be dissolved entirely."[2] after director Professor Peter Gøtzsche was forced to leave. Gotzsche stated that:
"Cochrane no longer lives up to its core values of collaboration, openness, transparency, accountability, democracy and keeping the drug industry at arm’s length." - Peter Gotzsche [3]
Without a transparent review process it is impossible to know which reviews to trust. FAIR reviews change that.
References
- Haddaway, N.R. and Westgate, M.J., 2019. Predicting the time needed for environmental systematic reviews and systematic maps. Conservation Biology, 33(2):434-443.
- Vesper, Inga (17 September 2018). "Mass resignation guts board of prestigious Cochrane Collaboration". Nature. doi:10.1038/d41586-018-06727-0.
- Peter C. Gøtzsche (November 8, 2018). "Peter C Gøtzsche: Cochrane—no longer a Collaboration". The British Medical Journal. Retrieved November 21, 2018.
